RNA-Seq, Microarray, and Multi-Omics Data Analysis Software
Subio Platform is data analysis software for RNA-Seq, microarray, and multi-omics data. It helps you visualize and interpret expression data and other omics measurements, and supports the integration of datasets across different omics types for exploratory analysis and biological interpretation.
See your data. Verify your results.
No black boxes. Just understanding.
No programming required. Works on Windows and Mac.
Got Results—but Not Sure What to Make of Them?
You followed the tutorial.
You ran the pipeline.
You got results.
But then you wonder—
- Is this normalization really appropriate?
- Does this cluster mean anything?
- Can these p-values be trusted?
Have you ever ended your analysis with these questions still unanswered?
R-based tools such as DESeq2 play an important role in RNA-Seq analysis.
However, beginner-oriented learning often emphasizes running commands and getting results first.
As a result, it is easy to follow commands step by step
without sufficiently examining data distributions or differences between samples.
GUI tools make it easy to obtain results.
However, they may not always make it easy to examine the assumptions behind the process
or the state of the data itself.
Subio Platform gives you a way to examine these questions visually.
It lets you analyze your data while actually seeing and understanding it.
Stop Running Tools. Start Understanding Your Data.
With Subio Platform, you can finally:
- Detect batch effects before they mislead your analysis
- Verify your results by examining the data behind them
- Combine visual workflows with R, Python, and AI
- Turn your data into a reusable research asset
Trusted by researchers since 2009.
See publications and user voices.
Most RNA-Seq tools run the workflow.
Subio Platform helps you understand it—visually.
See what’s really happening in your data—before you trust the results.
No programming required. Works on Windows and Mac.
Start exploring your data right away.
Start RNA-Seq Tutorial
No setup required. Includes demo data.
Is Running the Tool Enough to Solve the Real Problem?
Statistical tools such as DESeq2 and GSEA play an important role in RNA-Seq analysis.
However, obtaining statistical results is not the same as understanding, validating,
and applying those results in your research.
When analysis is completed with statistical tools alone,
modern omics analysis can face the following challenges.
Beyond Statistical Tools:
Four Challenges Often Encountered in Modern Omics Analysis
1. Lack of Visual Validation Risk of Missing Hidden Biases
Without visualization, batch effects and artifacts may go unnoticed.
2. Lack of Continuity Lost Knowledge and Poor Reproducibility
Can you reproduce or explain your analysis months later?
3. Low Efficiency Time Lost Outside Analysis
Too much time spent fixing environments and debugging scripts.
4. Lack of Exploration Static Reports Can End Exploration Too Early
Excel or PDF outputs can limit further exploration.
Comparison of RNA-Seq Analysis Approaches
Different tools offer different strengths and limitations.
The table below summarizes how they compare in usability, interpretability, and flexibility.
| Approach | Ease of Use | Interpretability | Flexibility |
|---|---|---|---|
| Command-line tools (R, Python) | Moderate | Moderate | High |
| GUI tools (especially browser-based) | High | Moderate | Limited |
| Subio Platform (GUI + script integration) | High | High | High |
Five Key Values Subio Platform Brings to Omics Analysis
1. See Your RNA-Seq Data Before You Trust the Results
Subio Platform provides a GUI designed specifically for omics data.
- Instantly visualize batch effects and biases
- Detect issues that may be less visible in some workflows
- Validate normalization and preprocessing visually
Why visualization matters
Visualization is an essential part of reliable analysis.
Batch effects are not uncommon, especially in large datasets.
See the relationship between clusters and batch effects at a glance.
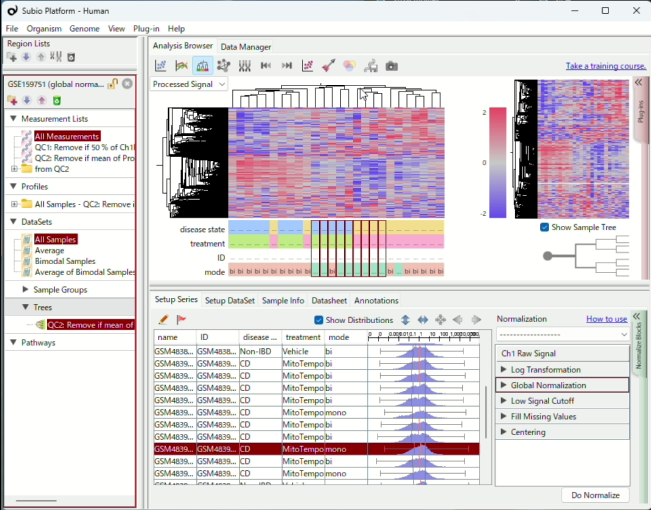
View the data distribution of each sample alongside your main plots.
With Subio Platform, you can visually verify your data at every step of the analysis.
Want to see a real example?
Even when using correction methods like ComBat,
you must verify whether the correction actually worked.
Without seeing raw data:
- You may mistake technical artifacts for biological signals
- You may trust “clean” normalized data that hides real problems
Good analysis starts with one simple rule:
A useful principle is to verify data visually before trusting it.
2. Combine Subio with R, Python, and AI
Subio Platform complements — not replaces — R and Python.
- Use Subio for intuitive visualization and data management
- Use R/Python for advanced statistical analysis
- Safely run AI-generated scripts in your local environment
Typical workflow
- Explore and prepare data in Subio
- Run analysis in R/Python (or AI-generated code)
- Import results back into Subio for validation and interpretation
For large datasets:
- Use Subio for exploratory analysis (e.g., ~100 samples)
- Then scale with scripts for full datasets
3. Turn Your Data into a Research Asset
Your data shouldn’t disappear after one analysis.
With Subio:
- Integrate your own data with GEO and TCGA datasets
- Reanalyze past experiments instantly
- Compare across platforms (RNA-Seq, microarray, etc.)
Your data becomes a living, searchable archive.
4. Use an Omics Analysis Environment Built to Last
Subio Platform itself is provided free of charge.
Data visualization and data management are available for free,
while more advanced analysis functions and support are provided through plug-ins and analysis services.
If you would like to see what you can do with the free core features, please also see:
What You Can Do with the Free Core Features of Subio Platform
For pricing details, please see the price list.
At the same time, Subio Platform is designed as an environment for long-term research use.
- Less affected by OS updates and changes in the analysis environment
- Helps reduce the risk of research projects being disrupted by changes in analysis tools
5. Transform Collaboration with SSA Files
Subio’s SSA format goes beyond static results.
It packages:
- Raw data
- Analysis results
- Parameters
- Notes and documents
into a single file.
This enables:
- Instant reproducibility
- Interactive reanalysis by collaborators
- A shift from “reporting” to discussion-driven science
What happens when you receive an SSA file?
This 90-second video shows how recipients can open an SSA file,
inspect the data, and start reanalysis without rebuilding the environment.
Why Visualization Matters
Visualization is not just a “nice-to-have” feature.
It is a critical process that determines whether your analysis is trustworthy.
Most tutorials focus on command-line operations.
But without visualization, you are effectively proceeding without fully understanding the overall structure of your data.
Batch effects are not a special case.
In omics data, they should be assumed to exist until proven otherwise.
The risk increases as the size of the dataset grows.
Even in small datasets, it cannot be assumed to be completely absent.
Correction methods such as ComBat will automatically produce results.
However, it is up to the analyst to verify whether the correction has actually worked.
If you inspect the raw data, you may recognize that a particular cluster is likely caused by batch effects.
But if you only analyze the corrected, superficially adjusted data,
you may mistakenly interpret such clusters as biologically meaningful.
This is very similar to what was often observed in microarray data analysis after RMA normalization.
(An example where data “cleaned” by RMA does not necessarily reflect the underlying biological reality is
here.)
Reliable analysis starts with a simple premise:
assume bias may exist—and verify it visually.
Explore Real-World Examples That Change How You Analyze Data
-
Apparent Differences That Can Appear After CPM Normalization
A real-data example of unexpected effects of normalization -
Before Taking FPKM/TPM Data as “Cleaned”
The importance of data validation, shown with GSE159751 -
Why Count Data Outperforms FPKM/TPM in RNA-Seq Analysis
Understand what actually improves reliability. -
Why You Need Multiple Visualizations — Not Just One
Avoid misleading conclusions from single plots.
How Subio Works with R, Python
In practice, data import, normalization, and preprocessing can all be handled within Subio Platform.
Filtering and standard statistical analyses (such as t-tests, PCA, and clustering) can be performed easily using Plug-ins.
For more advanced or specialized analyses, data can be exported as tab-delimited text and passed to R or Python,
with the results then re-imported into Subio Platform—maintaining a consistent analytical workflow.
As dataset size increases (as a rough guideline, around 100 samples or more),
the speed and scalability of command-line tools become increasingly valuable.
However, applying pipelines directly to the full dataset may lead to suboptimal or misleading results.
A more practical approach is to first analyze a subset of the data using Subio Platform’s visualization capabilities,
clarify your analysis strategy, and then implement it as a script to run across the full dataset.
How Subio Works with AI—Without Losing Control
Even without coding expertise, researchers can now generate R/Python scripts using AI.
However, determining whether the results are valid remains the responsibility of the analyst.
Subio Platform enables you to manage your data locally while integrating seamlessly with R and Python,
allowing the entire analysis workflow to remain within a local environment.
At the same time, it provides a framework to visually verify AI-generated results with your own eyes.
- [Preparation] Use Subio to understand the data and export datasets for scripting
- [Computation] Execute AI-generated code in R/Python
- [Interpretation] Bring results back into Subio for visual validation and interpretation
For an overview of this hybrid analysis approach, see the following article with practical examples:
For step-by-step examples, see:
What You Can Do with an Integrated Data Archive
Cross-dataset insights
Case Study 339: Gene expression is consistent across datasets. miRNA is not.
When multiple hepatocellular carcinoma (HCC) datasets were compared, gene expression remained consistent across platforms (RNA-Seq and microarrays), whereas miRNA expression showed striking discrepancies between datasets.
Importantly, miRNA expression appears internally consistent within each dataset. However, this issue cannot be recognized from a single dataset—it only becomes evident when multiple datasets are compared.
Case Study 189: Cross-study consistency builds confidence
When independent datasets on mouse mammary gland development were compared, highly consistent expression patterns were observed across studies.
This demonstrates that by accumulating data and enabling cross-dataset comparisons, it becomes possible to assess the reliability of information and identify results that can be trusted.
Scan Genes over Series tool (System Plug-in):
This feature enables searches similar to GEO Profiles, but applied to your own accumulated datasets within Subio Platform.
You can instantly explore how a gene of interest behaves across different datasets (Series),
transcending differences in species and experimental platforms.
What types of data can be analyzed and stored?
Subio Platform can import a wide range of omics data, including RNA-Seq, microarray, DNA methylation, proteomics, ChIP-Seq, and ATAC-Seq data. It not only helps you visualize and analyze these datasets, but also allows you to store them in a reusable form. You can also import public datasets from databases such as GEO and TCGA for re-analysis and comparative analysis.
- Table-based data (tab-delimited text files): RNA-Seq Gene Counts, TPM, FPKM, transcript-level or exon-level expression values, microarray signal values, DNA methylation ratios, proteomics abundances, and more
- Sequence data: RNA-Seq FASTQ files
- Genome browser formats: BED / WIG / BigWig (primary analysis results from ChIP-Seq, ATAC-Seq, and other assays, as well as signal data for genomic regions)
If you find it difficult:
Public databases contain a wide variety of omics data, and you may encounter data types that are unfamiliar.
Working with such data requires both broad knowledge and practical experience.
If you find certain datasets difficult to handle, you can use Subio’s Data Analysis Service.
We can process your requested datasets and deliver them in SSA format, ready to be added to your archive.
Why Long-Term Sustainability Matters
The sustainability challenge in bioinformatics tools
Bioinformatics has advanced greatly through free and open tools.
However, in academia, developing new tools is often valued more highly than maintaining existing ones.
As a result, many useful tools eventually become difficult to install, update, or use in changing computing environments.
From a user’s perspective, this can mean that a familiar tool may become unusable over time.
If that tool is central to a research workflow, the result can be delays—or even interruption—of a project.
A sustainable foundation through independent operation
Subio was established in 2008 as one approach to this challenge.
From the beginning, we focused on building an independently operated platform that could be maintained over the long term, without depending on project lifecycles, temporary funding, or policy changes.
Since then, we have continued to prioritize long-term development, maintenance, and support.
For developers and analysts: co-creation on the Subio Platform
Subio Platform can also serve as a foundation for creating and delivering new value through plug-in development and analysis services.
Tools can be distributed in parallel: as open-source software, and also as plug-ins within Subio Platform for users who need an easier and more integrated environment.
Learn more about plug-in development and distribution
Learn more about providing analysis services using Subio Platform
If you are interested in these opportunities, please feel free to contact us.
Why Sharing via SSA Files Matters
90-Second Demo: What Happens When You Receive an SSA File
Watch this short demo first to see how an SSA file lets collaborators open, explore, and reanalyze omics data immediately.
( View Content)
Even if raw data can be downloaded from public databases such as GEO,
only a limited number of researchers can actually reproduce published results
or reuse the data in their own studies.
On the other hand, receiving analysis results as static files such as PDFs or Excel sheets
rarely leads to meaningful biological discussion.
This situation—data exists, but cannot be explored—has long been a bottleneck in omics research.
SSA files address this problem by allowing collaborators to interact directly with the data
and reanalyze it from their own perspectives.
With SSA files, data sharing changes from one-way reporting into discussion-driven collaboration.
Why this matters beyond file sharing
Command-line tools are widely used not only because they are powerful,
but also because well-established learning environments exist around them.
At the same time, we still have much to learn about how to truly interpret omics data.
Analytical methods should ideally be refined through evaluation and feedback from experimental biologists.
In reality, however, such feedback often remains within the bioinformatics community,
where it may receive limited external validation or critical review from experimental researchers.
As AI makes it easier to incorporate advanced analytical methods,
it becomes even more important for researchers to examine results visually
instead of accepting them as black-box outputs.
If more experimental biologists can explore, question, and validate omics data directly,
real-world feedback can circulate more effectively.
We believe this kind of feedback loop can help bioinformatics evolve in a more practical,
reliable, and research-driven direction.
Who Subio Platform Is For
- Researchers who want to understand their data deeply
- Labs managing multiple datasets
- Scientists integrating public and private data
Who It’s NOT For
- Those focused only on programming skills
- One-off analyses with no need for reuse
- Fully automated pipelines without interpretation
Support & Services
Frequently Asked Questions about RNA-Seq Data Analysis
These are common challenges researchers face when using standard RNA-Seq tools.
Why do many RNA-Seq tools fail to detect batch effects?
Many RNA-Seq tools focus on executing pipelines, and may provide limited support for visual inspection. As a result, batch effects and technical biases can remain hidden.
Subio Platform allows you to visualize data distributions and sample relationships at every step, making it easier to detect issues before they affect your results.
Why is it difficult to interpret RNA-Seq results with standard tools?
Standard tools often produce statistical outputs without providing sufficient context for interpretation. This can make it hard to determine whether results reflect true biological signals or artifacts.
Subio Platform helps you interpret results by combining statistical outputs with visual exploration, so you can understand what is happening in your data.
What are the limitations of GUI-based tools?
While GUI tools improve usability, many are optimized for ease of use, which may limit deeper analysis in some cases. They may limit flexibility or prevent detailed validation of intermediate steps.
Subio Platform combines an intuitive GUI with the ability to integrate R, Python, and AI workflows, allowing both ease of use and analytical depth.
Master Analysis, Not the Tool.
Do you want to learn commands?
Or do you want to learn data analysis?
The choice is yours—right now.
Why not see for yourself, using your own data?
Download Free
Plug-ins are 30% off for first-time users,
and can be discounted up to 50% under certain conditions
See all discount options and conditions
To download Subio Platform, enter your information below, agree to the terms and conditions, and click the download button.
System Requirements
- Windows 10 or 11 (64-bit) / macOS 10.14 Mojave or later
- 2 GHz or faster processor
- 4 GB or more of RAM
- 5 GB or more of available hard-disk space
- 1024 x 768 display
Next Step
Thank you for downloading.
Start by loading your data into Subio Platform
and visually checking distributions and potential batch effects.
If you are new, you can begin with a tutorial using demo data:
RNA-Seq Tutorial
How to Move Forward with Your Analysis
With Subio Platform,
data visualization and management are fully available for free.
Core analysis steps (such as statistical analysis, clustering, and differential expression analysis)
can also be performed using R or Python at no cost.
However, this often requires writing and adjusting code, which can take significant time and effort.
By using plug-ins,
you can perform these analyses more efficiently,
without spending time on operations—
allowing you to focus on understanding your data and interpreting the results.
Some functions are also available only through plug-ins,
and can be difficult to reproduce using scripts alone.
Plug-ins are affordably priced,
and you can start with a 5-day free trial.
Results generated using plug-ins remain fully accessible,
even after the plug-in license has expired—
your data and results are always yours.
Plug-in Discounts
Plug-ins and plug-in bundles are 30% off for first-time users.
In addition,
You can receive $50 or €40 off per referral link
(based on your billing currency, up to 50% total discount)
* To apply, publish your link before purchase
and include the URL when requesting a quote.
* Available for both new and existing users.
Where Would You Like to Start?
-
Learn the workflow step by step and explore at your own pace
RNA-Seq Analysis Tutorial -
Move forward with your own data efficiently while building understanding
Data Analysis Service
Online Training